Live Cell Super-Resolution Imaging Research Team
Team Leader
Katsumi Midorikawa
D.Eng.

Contact
kmidori [at] riken.jp
Live Cell Super-Resolution Imaging Research Team,
RIKEN Center for Advanced Photonics
#S411 4F Bioscience Building,
2-1 Hirosawa, Wako, Saitama 351-0198 Japan
Related links
Laboratory Website
Live Cell Super-Resolution Imaging Research Team![]()
Laboratory on RIKEN Website
Live Cell Super-Resolution Imaging Research Team | RIKEN![]()
Outline
Light is a cutting-edge tool for life science research. Development of useful fluorescent probes and the advancement of light microscope technologies have brought us a new world of “live” imaging within a cell. We are developing super-resolution confocal live imaging microscopy (SCLIM) by the combination of a high-speed confocal scanner and a high-sensitivity camera system. With this method we will observe membrane trafficking and organellar dynamics in living cells at high-speed and sub-wavelength space resolution (4D) and elucidate underlining molecular mechanisms. We will also try to extend this technology to medical and pharmacological applications.
Fields
Biological Sciences, Complex systems, Interdisciplinary science and engineering, Biology / Biochemistry
Keywords
membrane traffic, vesicular transport, super-resolution live imaging, confocal microscopy, organelles
Subjects
- Development of super-resolution live imaging microscopy
- Molecular mechanisms of intracellular membrane trafficking
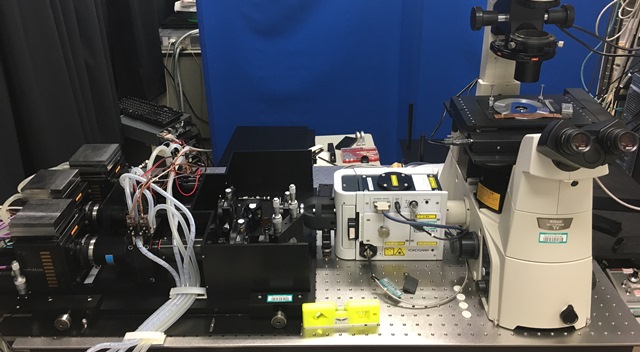

SCLIM2 microscope
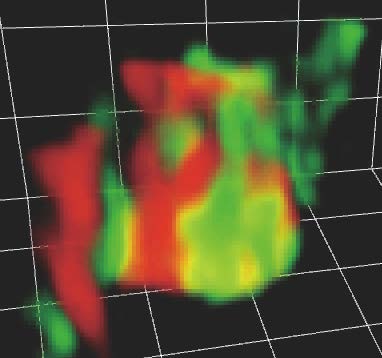
Yeast trans-Golgi network (red) and clathrin-coated vesicles (green) assembling and leaving there. Captured from a high-speed 3D movie taken by SCLIM2.
Selected Publications
- Nakano, A.: “The Golgi apparatus and its next-door neighbors” , Frontiers Cell Dev. Biol.10:884360 (2022).
- Shimizu, Y., Takagi, J., Ito, E., Ito, Y., Ebine, K., Komatsu, Y., Goto, Y., Sato M., Toyooka, K., Ueda, T., Kurokawa, K., Uemura, T., and Nakano, A.: “Cargo sorting zones in the trans-Golgi network visualized by super-resolution confocal live imaging microscopy in plants” , Nat. Commun. 12:1901 (2021).
- Rodriguez-Gallardo, S., Kurokawa, K., Sabido-Bozo, S., Cortes-Gomez, A., Ikeda, A., Zoni, V., Aguilera-Romero, A., Perez-Linero, A. M., Lopez, S., Waga, M., Araki, M., Nakano, M., Riezman, H., Funato, K., Vanni, S., Nakano, A., and Muñiz, M.: “Ceramide chain length-dependent protein sorting into selective endoplasmic reticulum exit sites” , Sci. Adv. 6:eaba8237 (2020).
- Tojima, T., Suda, Y., Ishii, M., Kurokawa, K., and Nakano, A.: “Spatiotemporal dissection of the trans-Golgi network in budding yeast” , J. Cell Sci. 132:jcs231159 (2019).
- Kurokawa, K., Osakada, H., Kojidani, T., Suda, Y., Asakawa, H., Haraguchi, T., and Nakano, A.: “Visualization of secretory cargo transport within the Golgi apparatus in living yeast cells” , J. Cell Biol. 218:1602-1618 (2019).
Publications
Members
| Katsumi Midorikawa | Team Leader |
| Akihiko Nakano | Dupty Team Leader |
| Kazuo Kurokawa | Senior Research Scientist |
| Takuro Tojima | Senior Research Scientist |
| Kumi Matsuura | Research Scientist |
| Natsuko Jin | Research Scientist |
| Daisuke Miyashiro | Technical Scientist |
| Tomohiro Uemura | Visiting Scientist |
| Yasuyuki Suda | Visiting Scientist |
| Yoko Ito | Visiting Scientist |
| Emi Ito | Visiting Scientist |
| Kumiko Ishii | Technical Staff Ⅱ |
| Miho Waga | Technical Staff Ⅱ |
| Hideo Hirukawa | Technical Staff Ⅱ |

